| bioLegato Online Help |
Open
and Import Files
The
bioLegato File --> Open
menu is "hardwired"
into bioLegato, and can open the main file types that are
intrinsic to
the bioLegato program: GDE (.gde), GenBank (.gen), FASTA
(.fsa) and
flat (.flat).
Each bioLegato
interface also has
one or more Import menus, which have been programmed for
data types
that are specific to each bioLegato interface.
The Open Menu
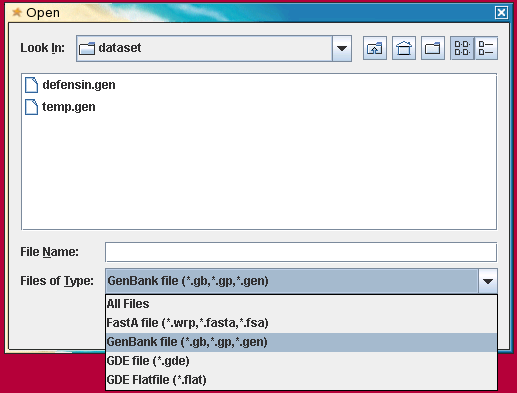 The
File --> Open menu
will only
read files in GDE (.gde), FASTA (.fsa), GenBank
(.gen), or flatfile (.flat) format.
The
File --> Open menu
will only
read files in GDE (.gde), FASTA (.fsa), GenBank
(.gen), or flatfile (.flat) format. GenBank format - bioLegato can read any file in GenBank format, as defined in the GenBank release notes from NCBI [ ftp://ftp.ncbi.nih.gov/genbank/gbrel.txt]. Files originally-read in GenBank format can also be saved as GenBank files, because biolegato stores the annotation portion of each GenBank entry as text, unchanged from the original file.
One word of caution is in order regarding saving in GenBank format. If a sequence was read in a different format (eg. FASTA) bioLegato will save in a pseudo-GenBank format, in which only minimal annotation fields are written, since the data for complete annotation does not exist in most other file formats. using File --> SaveAs. However, this format loses some of the annotation information from the original GenBank file, and in other cases, rearranges data, so that GenBank files generated by bioLegato do NOT comply with the official standard.
Examples:
thionin.gen
- file containing two thionin sequences directly retrieved
from NCBI.
thionin.biolegato.gen - thionin.gen was read into bioLegato and then saved using SaveAs.
thionin.biolegato.gen - thionin.gen was read into bioLegato and then saved using SaveAs.
| Warning:
Although the differences between these two formats may
look minor, they
may break programs that read GenBank files. Therefore,
when retrieving
GenBank files using biolegato or dbiolegato, it is
best to save them
directly to a
file, rather than sending them to biolegato and later
saving. |
GDE format - This format is similar to the ASN.1 format used in databases. It preserves most of the annotation from the original GenBank file.
Example:
thionin.gde
- thionin.gen was read into biolegato and then saved using
SaveAs.
GDE flatfile format - This is a minimal format, which only retains the name and sequence data. It is used by biolegato to create temporary files, and there is no way to directly save to this format.
Example:
thionin.flat
- bioLegato flat file using thionin sequences from above.
The name of each "sequence" in a GDE flat file indicates the type of data:
# -
DNA/RNA
% - protein
@ - mask sequence
" - text
% - protein
@ - mask sequence
" - text
For example, a bioLegato flat file containing a list of GI numbers for DNA sequences from Brassica insularis (TaxID 69183) might look like this:
"69183.dna.gi
75707982,57638973,57638879,44888690,9719214,2598264,2598263
bldna,blprotein,blnalign,
blpalign
 The File
--> Import
Foreign Format menu calls readseq,
which can
translate a number of sequence formats into a pseudo-GenBank
format.
The ouput is then read into biolegato. It is important to
realize that
no
other file format contains the complete information found in
GenBank
files. Usually, only the name of the sequence and the sequence
itself
are retained. Even if this function is used to read in a GenBank
file,
the fact that it is processed by both readseq and biolegato
means that
the
full annotation may not be retained.
The File
--> Import
Foreign Format menu calls readseq,
which can
translate a number of sequence formats into a pseudo-GenBank
format.
The ouput is then read into biolegato. It is important to
realize that
no
other file format contains the complete information found in
GenBank
files. Usually, only the name of the sequence and the sequence
itself
are retained. Even if this function is used to read in a GenBank
file,
the fact that it is processed by both readseq and biolegato
means that
the
full annotation may not be retained.| Note
on circular sequences: The only common sequence
file format that
can
specify whether a sequence is circular or linear is
GenBank. Always use
this format when working with circular sequences. |
SaveAs
| The SaveAs menu can be used
to
save in any of the four formats supported by biolegato. |
 |
Export Foreign Format
| The File --> Export Foreign
Format
menu also calls readseq,
to create files in numerous formats. Again, since
biolegato can not
necessarily export all of the original annotation, some
annotation my
not appear in the exported files. Example: |
 |
bltable
Todo: add documentation for bltable
blmarker
| blmarker works with
discrete state data, such as molecular markers. The File --> Import
Phylip Discrete Data menu translates a Phylip
discrete data file
into biolegato flat file, and imports the flat file. Example: Similarly, the File --> Export Phylip Discrete Data menu saves discrete state data as a flat file, which is then translated into a Phylip discrete data file. |
  |
Importing Data from a Spreadsheet
Molecular marker data is often assembled in a spreadsheet, as illustrated below. Marker data for each species, isolate, or strain should be in a row. The first column is holds tha name of the species, isolate or strain, and the remaining columns have the marker data, such that each column is a different marker (ie. locus).
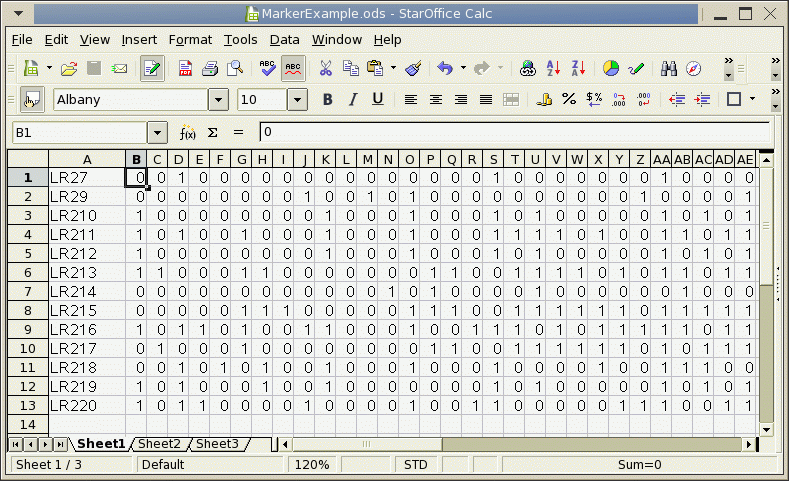
To use this data in blmarker, it must be saved as a Comma-Separated Value file (.csv).
If you are using StarOffice or OpenOffice Calc,
- choose File --> SaveAs
- set the File Type to "Text CSV (.csv)".
- Make sure the field delimiter is a comma (,).
- choose File --> SaveAs,
- Other formats
- set Save as type to "CSV (Comma delimited) (*.csv)".
For example, if your spreadsheet file was MarkerExample.xls, then save your data to MarkerExample.csv.
At this point it is a good idea to check your .csv file by looking at it in a text editor. It should look something like this:
"LR27 ",0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,1,0,1,1,1,1,0,0,0,1,1,1,1,1,1,1
"LR29 ",0,0,0,0,0,0,0,0,1,0,0,1,0,1,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,1,0,0,0,1,0,0,0,0,0,0,1,1,1,1,1,1,1,1,1,1,1,0,0,1,1,1,1,1,1,1
"LR210 ",1,0,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,1,0,1,0,0,0,0,0,1,0,1,0,1,1,0,0,1,0,0,0,0,0,0,1,1,0,1,0,1,1,1,1,0,1,0,0,1,0,0,1,1,1,1
"LR211 ",1,0,1,0,0,1,0,0,0,1,0,0,0,1,0,0,0,1,1,1,1,0,1,0,0,1,1,0,1,1,1,0,0,1,0,0,0,0,0,0,1,0,0,1,1,1,1,0,1,0,1,1,1,1,0,1,1,1,1,1
"LR212 ",1,0,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,1,0,1,0,0,0,0,0,1,0,1,0,1,1,0,0,1,0,0,0,0,0,0,1,0,0,1,0,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1
"LR213 ",1,1,0,0,0,1,1,0,0,0,0,0,0,0,1,1,0,0,1,1,1,1,0,1,0,1,1,0,1,1,1,1,1,1,0,1,1,1,1,1,1,1,0,1,0,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1
"LR214 ",0,0,0,0,0,0,0,0,0,0,0,0,1,0,1,0,0,0,0,1,0,0,0,0,0,0,1,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,1,0,0,0,0,1,0,0,1,0,0,1,1,1,0,0,0,0
"LR215 ",0,0,0,0,0,1,1,1,0,0,0,0,0,1,1,1,0,0,1,1,1,1,1,1,0,1,1,1,1,1,1,0,1,1,1,0,1,1,1,1,1,1,0,1,0,1,1,0,1,1,1,1,1,1,1,1,1,1,1,1
"LR216 ",1,0,1,1,0,1,0,0,1,1,0,0,0,1,0,0,1,1,1,0,1,0,1,1,1,1,1,0,1,1,1,1,0,1,0,0,0,0,0,0,1,1,1,1,1,1,1,1,0,1,1,1,1,1,1,1,1,1,1,1
"LR217 ",0,1,0,0,0,1,0,0,0,0,0,0,0,0,1,1,0,0,1,1,1,1,1,1,0,1,0,1,1,1,0,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1
"LR218 ",0,0,1,0,1,0,1,0,0,1,0,0,0,1,0,0,0,1,0,0,0,0,1,0,0,1,1,1,0,0,0,1,0,1,0,0,0,0,0,0,0,1,1,0,0,1,1,1,0,1,1,1,1,1,1,1,1,1,1,1
"LR219 ",1,0,1,0,0,0,1,0,0,1,0,0,0,0,0,0,0,1,0,1,0,0,0,0,0,1,0,1,0,1,1,1,0,0,0,0,0,0,0,0,0,1,1,0,0,1,1,0,1,0,1,1,1,1,1,1,1,1,1,1
"LR220 ",1,0,1,1,0,0,0,0,1,0,0,0,0,1,0,0,1,1,0,0,0,0,0,1,1,1,0,0,1,1,1,1,0,1,0,0,0,0,0,0,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1
Note: The example above shows that the name field is enclosed in quotes. These are okay, as quotes will be automatically stripped out when the file is read by blmarker.
| To import this file
into blmarker, choose File
-->
Import
Discrete Data from CSV file. The .csv file is read into blmarker, which should appear as shown below. Note: you could also convert a .csv file into a Phylip file at the Unix command line using csv2phyl.sh. |
 |