TUTORIAL: Genome Assembly |
December 1, 2023 |
TUTORIAL: Genome Assembly |
December 1, 2023 |
| A
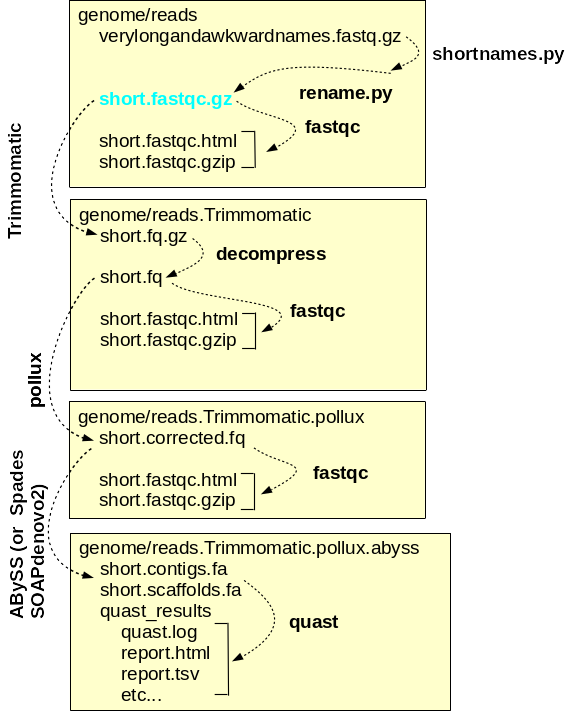
simplified workflow for genome assembly is shown at right.
Genome assembly is carried out within the genome directory, and for each major step, a separate subdirectory is used. By convention, the names of subdirectories tell the series of programs used to generate the results in each directory. Raw read files are saved in the raw directory, and symbolic links to these files, with short, meaningful names, are created. Sequencing adaptors are removed from theTrimmomatic raw reads by and the files containing the trimmed reads are saved in reads.Trimmomatic. The trimmed reads are used as input by pollux, which corrects errors in the reads and writes the corrected reads to the pollux directory. At each step, FASTQC is run to check the properties of the reads. Finally, the assembly itself is done using programs such as ABySS or Spades or SOAPdenovo2, which produce contig files and scaffold files. For each assembly, quast produces an reports with extensive statistics that can guide the choice of which assembly is best, or which assemblies should be repeated with different a parameters for improvement. |
 |