TUTORIAL:
Genome
Assembly
|
June 10, 2019 |
TUTORIAL:
Genome
Assembly
|
June 10, 2019 |
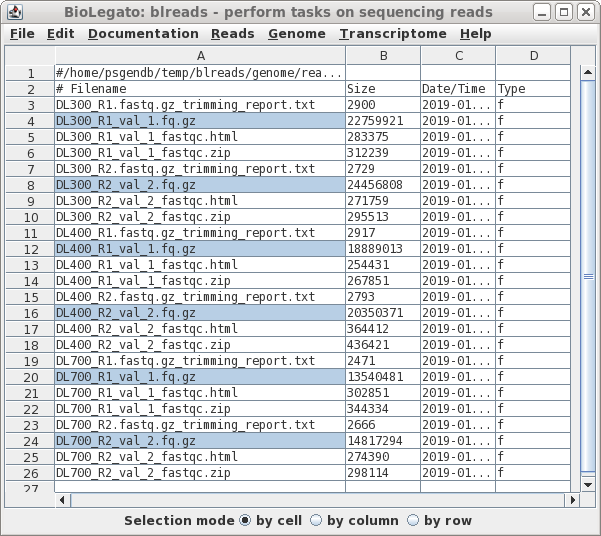
| OPTIONAL: HOW TO
DECOMPRESS GZIPPED FILES In the previous step, the instructions said to set compress output files using gzip to No. If, however, you did have trimmed reads in compressed form, blreads has an uncompress function. If your reads are not compressed, skip to Running Pollux below. Unfortunately, Pollux can't read compressed fastq files, so we have to decompress these files. First, select all compressed fastq.gz files. Next, choose File --> Decompress and click on Run. On large files, Decompress may take a few minutes. When complete, the window will be refreshed, showing that the .gz extension is gone from the fastq (.fq) files. |
 |
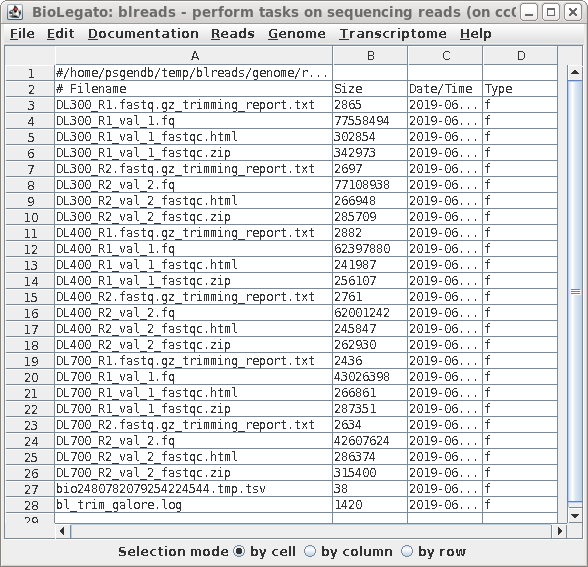
| Running pollux Your blreads window should now appear as shown at right. The read files will have the .fq file extension. |
 |
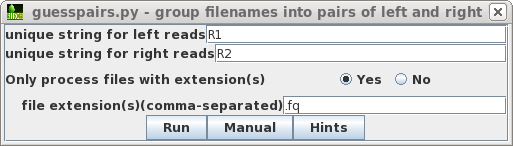
| We only want to process read files, and not
the other files in the directory. For this dataset, all read
files use the .fq extension. In some cases, you may need to
specify more than one set of file extensions as a
comma-separated list eg. .fq,.fastq Clicking on the Hints
button will give a more detailed explanation of these
parameters. |
 |
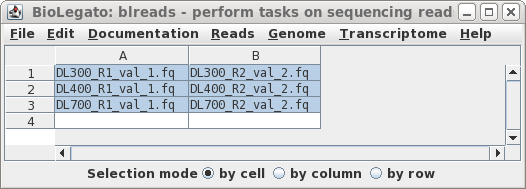
| Clicking on Run will bring up a new
blreads window with the best quess of file pairing, in two
columns. Usually, guesspairs.py gets it right. Files for
which a pair cannot be found (ie. single-end reads) would be
listed in a single column. To run Pollux with these read pairs, choose Edit --> SelectAll, and then Reads --> Pollux. |
 |
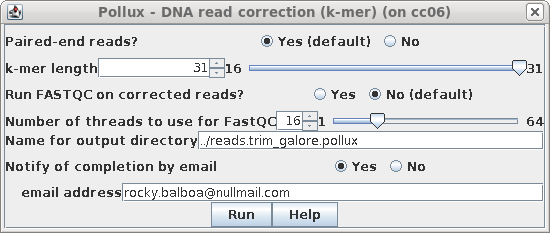
| By default, pollux will write corrected reads
to a directory in the parent directory called
reads.trimmed.pollux. To be more precise, let's call the
output directory ../reads.trim_galore.pollux Pollux may take awhile to run, so you may wish to set "Notify of completion by email" to Yes and type in an email address. |
 |
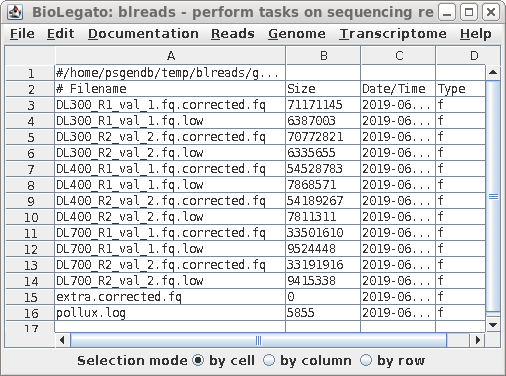
| The output is written to the
../reads.trim_galore.pollux directory, whose contents is
shown at right. The report in pollux.log summarizes the numbers of errors corrected of different types eg. insertions, deletions, corrections, homopolymer corrections. The high quality corrected reads are found in files with the .corrected.fq extension. Corrections based on low k-mer counts have the .low extension, and are generally not used. |
 |