As with so many words in the sciences, the suffix -ploid is derived from a Greek word meaning fold. When we talk about ploidy, we're describing the increase or decrease of genetic material by whole chromosome sets. Thus, when we say "wheat is hexaploid", we mean that it has six (yes, six!) full chromosomal sets. In fact, as long as the increase or decrease is in fact by another full set, we can say that an organism or species is euploid (true-fold), meaning that individual has one complete chromosome set or multiples of the basic number of chromosomes characteristic of the species. Sometimes the multiple isn't a whole number - that is, sometimes only part of a set is added, which is what we call aneuploid (not-true-fold).
Another concept we need to discuss is the N and X concepts. N may already be familiar to you, as the gametic number of chromosomes (akin to the complexity of the genome). We can say that cells are N (haploid complement) or 2N (diploid complement). X, on the other hand, is more relevant when discussing how species are related to each other. X is the basic number of chromosomes, with respect to a common ancestral species (ie. x is the "original" set of chromosomes, from which contemporary species are derived).
We can classify all organisms with a ploidy level higher than two (ie. triploids, tetraploids, pentaploids, etc.) as polyploids more generally. Polyploids are prominent in the plant kingdom, making up 30-35% of angiosperm species, and 75% of the Graminae (grass) species. Polyploids have advantages in a wider ecological range of tolerances. In seed plants, the endosperm is normally triploid. Therefore, we can infer that seed plants have intrinsic mechanisms for tolerating triploidy. Polyploids are less common in animal species, particularly among mammals. However, polyploidy is often seen in amphibians and salmonid fishes.
In addition to specifying the exact ploidy level, we can also
categorize polyploids by where they are getting their extra
chromosome sets. To do this, we use the prefixes auto
(self), and allo (other).
Autopolyploids are individuals with chromosome sets characteristic of the species. The chromosome sets are homologous to each other and pair fully at meiosis. Autotriploidy 3x (AAA), autotetraploidy 4x (AAAA), autopentaploidy 5x (AAAAA), autohexaploidy 6x (AAAAAA) represents 3, 4, 5, 6 homologous basic genomes per cell.
Allopolyploids are individuals that arise after natural or experimental crossing of 2 or more closely-related species or genera and contain the chromosome sets of the parents. Two parental genomes are derived from a common ancestor, but have diverged over time through mutational events. Thus the genomes have enough sequence similarity to pair, but pairing may be imperfect. Chromosome pairing at meiosis is limited to bivalent pairing. The entire genome of parental species A is sufficiently diverged from that of parental species B to prevent pairing. This suggests that the 'homology searching' mechanism that acts during synapsis requires some threshold level of similarity before pairing can occur. An initial cross between A and B followed by chromosome doubling produces a functional diploid with two new AB sets AABB. Wheat is in fact an allohexaploid, having the genome set AABBDD.
Also possible is a combination of the two, or a autoallopolyploid: an individual with basic genome originating from both types of polyploidization.
Autopolyploids differ in phenotype "quantitatively" from their parents. That is, the only difference is in ploidy level, or number of chromosome sets, not in kinds of chromosomes. In plant breeding, autopolyploidy is encouraged on purpose, to achieve these benefits:
The disadvantages lie in:
However, some crops such as triploid potatoes are easily reproduced vegetatively and are therefore very competitive agronomically.
Autotriploids possess three sets of homologous chromosomes in their cells. They arise from the fertilization of an unreduced egg (2N) by a normally reduced sperm (N) of an originally diploid species. Autotriploidy can arise spontaneously, and can also be induced. Autotriploids have been identified in at least 68 different plant genera. They are phenotypically more vigorous but are not competitive at reproduction due to a high level of ovule abortion. This is advantageous in banana and watermelon where seedless varieties are preferred.
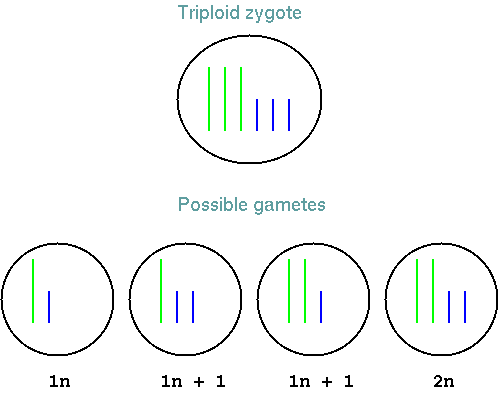
The problem with triploids comes during (of course) meiosis.
Triploid zygotes can produce many possible gametes, because each
gamete can get either one or two copies of each chromsome, in many
possible combinations. Consider a species with only two
chromosomes. Even in this simple case, only one out of four
possible gametes will have the normal haploid complement of
chromosomes. Aneuploid gametes are usually deleterious. In a
mating with a normal haploid gamete, the tetraploid gamete would
produce a triploid, while a 2N + 2N mating would give a
tetraploid, which may or may not be deleterious.

The number of possible combinations of chromosomes increases rapidly as the number of chromosomes increases. Therefore, the number of balanced gametes drops rapidly for genomes with larger numbers of chromosomes.
| To start, let's review what normal pairing looks like: | 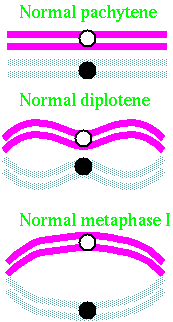 |
|
Autotriploidy is comparable to multiple primary trisomics. The three homologous chromosomes can form trivalents as well as bivalents with univalents. Trivalent association is more common with longer chromosomes. Any region can pair, but only two chromosomes can pair in any one region. If two of three homologues pair along their entire length, trivalents cannot form because one chromosome will be left completely unpaired, resulting in a bivalent and a univalent. If all three chromosomes are involved, chiasma formation may result in paired segments.
Figure 6.20. A, possible trivalent configurations at diakinesis in an autotriploid, based on crossing over and pachytene chromosome pairing.(Redrawn from Kuspira et al., 1986. Can. J. Genet. Cytol.28:867-887.) |
Autotetraploids can appear spontaneously in plants as a result
of non-disjunction in diploids either in the meristematic tissue
or somatic cells. They can also originate in reproductive tissue
through the formation of unreduced gametes. They can be
artifically produced through the application of colchicine or
nitrous oxide. Colchicine is the most effective in the largest
number of plant and animal species. Doubling occurs because the
spindle fibre formation is disrupted preventing the chromosomes
from migrating to the opposite poles. In general autotetraploids
are slow in growth with large dark green leaves. In some species,
autotetraploids are larger than the diploid with larger cell size.
In self-pollinating species, the autotetraploids are smaller or
similar to the diploids. Each chromosome is present four times and
theoretically the affinity is equal between all four sets of
chromosomes. There are ten possible quadrivalent configuration of
the four homologous chromosomes with any other combination of
univalents, bivalents, and trivalents. However, mostly chains
(convergent-a; parallel-a) and rings (convergent-b; parallel-b)
were observed. Most of the time, either all 4 homologues try to
pair, resulting in quadrivalents, or 2 sets of 2 chromosomes try
to pair, resulting in 2 sets of bivalents.
In the diagram below, note that all pairings have the full set of four chromosomes. The only difference is in which ends remain paired at metaphase I.
Any mutation begins as a single molecular event. Its frequency in a diploid population of size P is therefore 1/P. In other words, out of P individuals in the population, only 1 is polyploid as the result of the initial chromosome doubling event. To establish a new population in which all individuals have the new trait requires selfing or matings betwen closely-related relatives. The mode of reproduction can have a great influence on whether or not polyploidies can become established. For example, parthenogens - organisms that use asexual reproduction in which ovum develops into a zygote without fertilization - may have a different incidence of polyploidy.
| Taxa | Reproductive Mode | 2x/3x | 3x | 4x | Other |
|---|---|---|---|---|---|
| Insects | Parthenogens | 13 | 34 | 16 | 16 |
| Sexual | 0 | [1] | 0 | [2] | |
| Fish | Parthenogens | 1 | 6 | 0 | 2 |
| Sexual | 0 | 0 | 10 [1] | 12 | |
| ? | 0 | 2 | 12 [1] | 13 | |
| Amphibians | Parthenogen | 1 | 0 | 0 | 2 |
| Sexual | 0 | 0 | 18 [2] | 6 | |
| ? | 0 | 0 | 0 | 1 | |
| Reptiles | Parthenogens | 3 | 11 [1] | 0 | 0 |
| Sexual | 0 | 0 | 0 | 1 | |
| Mammals | Sexuals | 0 | 0 | 1 | 0 |
| Otto SP, Whitton J (2000) Polyploid incidence
and evolution. Ann. Rev. Genet. 34:401-437. This table summarizes Web Table 1, ignoring cases where polyploids are rare. Polyploid species are grouped by ploidy level and mode of reproduction, where '?' denotes that the reproductive mode is unknown to the authors. Values in brackets refer to additional cases where there is doubt about the polyploid nature of the species. |
|||||
The bias toward polyploidization in parthogens, compared to sexually-reproducing animals, is striking. Parthogens do not require fertilization by another individual, eliminating most of the aneuploids that would other wise result from a mating between haploid and polyploid gametes. In some cases, the ovum may require stimulation by sperm, although fertilization per se does not take place. In essence, parthenogenesis amounts to cloning. Parthenogenesis is distinct from hermaphrodism, in which an adult has both male and female sex organs, both of which undergo meiosis to produce gametes, allowing self-fertilization.
In plants, species are capable of selfing, which increases the
likelihood that two polyploid gametes will contribute to a zygote,
compared to outcrossing species. Once a population of polyploids
are established, matings with other polyploids are more likely to
give viable progeny than matings with normal diploids. This is the
beginning of reproductive isolation, leading to speciation.
| An allopolyploid is an individual dervied from interspecific hybridization from two or more genomically distinct and distantly related diploid species, followed by chromosome doubling of the sterile F1. Due to the highly divergent genomes, the chromosomes lack sufficient homology to form the tight bivalents required for equal segregation. Normal chromosome pairing and seed fertility are restored by doubling the chromosomes to a condition known as amphiploidy. In allopolyploid or amphiploid, multivalents are not formed in meiosis. Wheat, oats, canola are three agronomically important crops which are polyploid. | 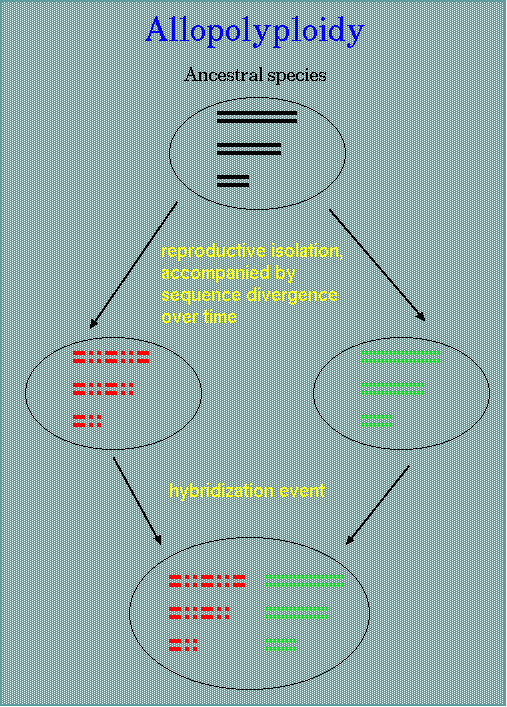 |
It's important to emphasize that the two superficially foreign genomes must have recently diverged from a single common ancestor. Allopolyploidy really should be thought of as a hybridization between two recently-diverges species, which have very closely-related genomes.
Aside from major chromosomal rearrangements, each chromosome in
one donor species has a counterpart in the other donor species.
Because both chromosomes are derived from the same ancestral
chromosome, they are said to be homeologous.
Homeologous chromosomes may or may not be able to pair in meiosis.
In wheat, there are loci known to inhibit pairing of homeologous
chromosomes. The origin of the allopolyploid species has been
demonstrated by crossing the diploid species and doubling the
chromosome numbers of the hybrid plant (synthetic). The
artificially produced allopolyploid is then crossed to the
tetraploid species with the corresponding chromosome number. The
homology of the chromosomes is then observed through the number of
bivalents formed at meiosis. If the homeologous chromosomes all
pair, one from the synthetic allopolyploid, and one from the
tetraploid found in nature, we can conclude that the tetraploid
originated from the two diploids that were used to create the
synthetic allopolyploid. This has been done, for example, with
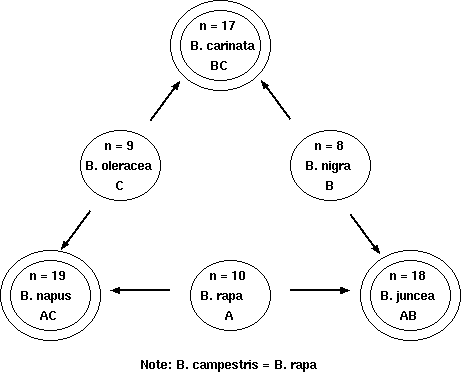
synthetic allotetraploids made from diploid Brassicas.
Allotetraploidy in Brassica spp. Relationships among diploid and allotetraploid Brassica. from U, N. 1935, Jpn. J. Bot. 7:389-452 |
 |
Allohexaploidy in wheatIn order to find out which of the wild species are the progenitors of wheat, geneticists have looked at geographical distribution, morphological features, karyotypes, C- and N-banding patterns, meiotic chromosome pairing, nuclear DNA content, in situ hybridization, restriction fragment patterns of rRNA, DNA hybridization, and so on. |
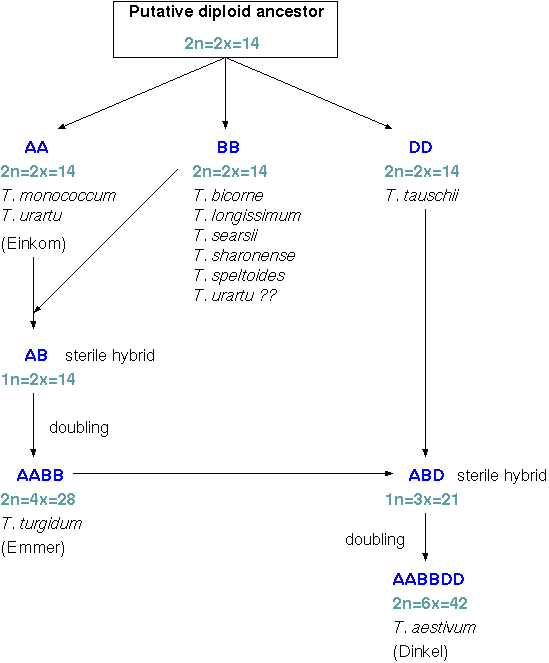 |
The genus Triticale was born out of the combination
of wheat (Triticum) and rye (Secale). It exists
at several ploidy levels, and the hexaploid and octoploid Triticales
have recieved the greatest amount of breeding effort.
Hexaploid triticale (primary) is obtained after doubling the chromosomes of the F1 hybrid between tetraploid wheat (AABB) and diploid rye (RR). The triploid F1 hybrid by itself would not be fertile. At meiosis, chromosomes would have no homologues with which to pair. Doubling the chromosomes results in normal meioses, since all chromosomes can now pair.
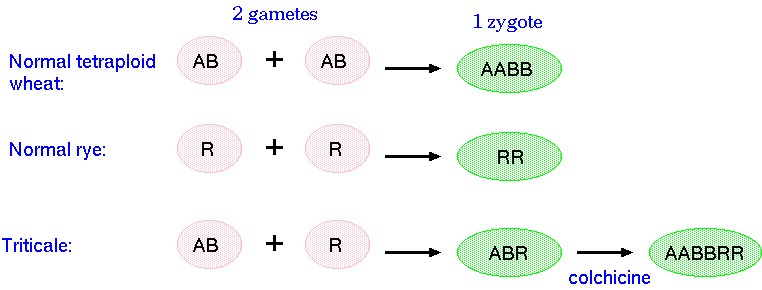
The genome formula for hexaploid triticale is 2n = 6x = 42 and its genome composition is AABBRR. Octoploid triticale (secondary) was obtained by crossing hexaploid wheat (AABBDD) to rye. The genome formula for octoploid triticale is 2n=8x=56 and its genome composition is AABBDDRR.
Problems with hexaploid hybrids included cytogenetic instability, reduced seed set and shrivelled seed which were major constraints for successful productivity. The cytological instability was attributed to cytoplasmic effects, differences between duration of meiosis and DNA content of wheat and rye, and interaction between wheat and rye chromosomes.
In octoploid triticale, a lower degree of aneuploidy, more stable meiosis, and higher fertility are seen, compared to hexaploid triticale. They are characterized by good winter hardiness, high protein content,good baking quality, early flowering, seed maturity and large seed size. However, seed sterility, disease (eg. ergot) and early sprouting are still problematic.